Attention
This is a new revised release, now using data from E. coli to speed the analysis steps up. The former tutorial based on fungi data can be accessed at https://genomics-fungi.sschmeier.com/.
Computational Genomics Tutorial¶
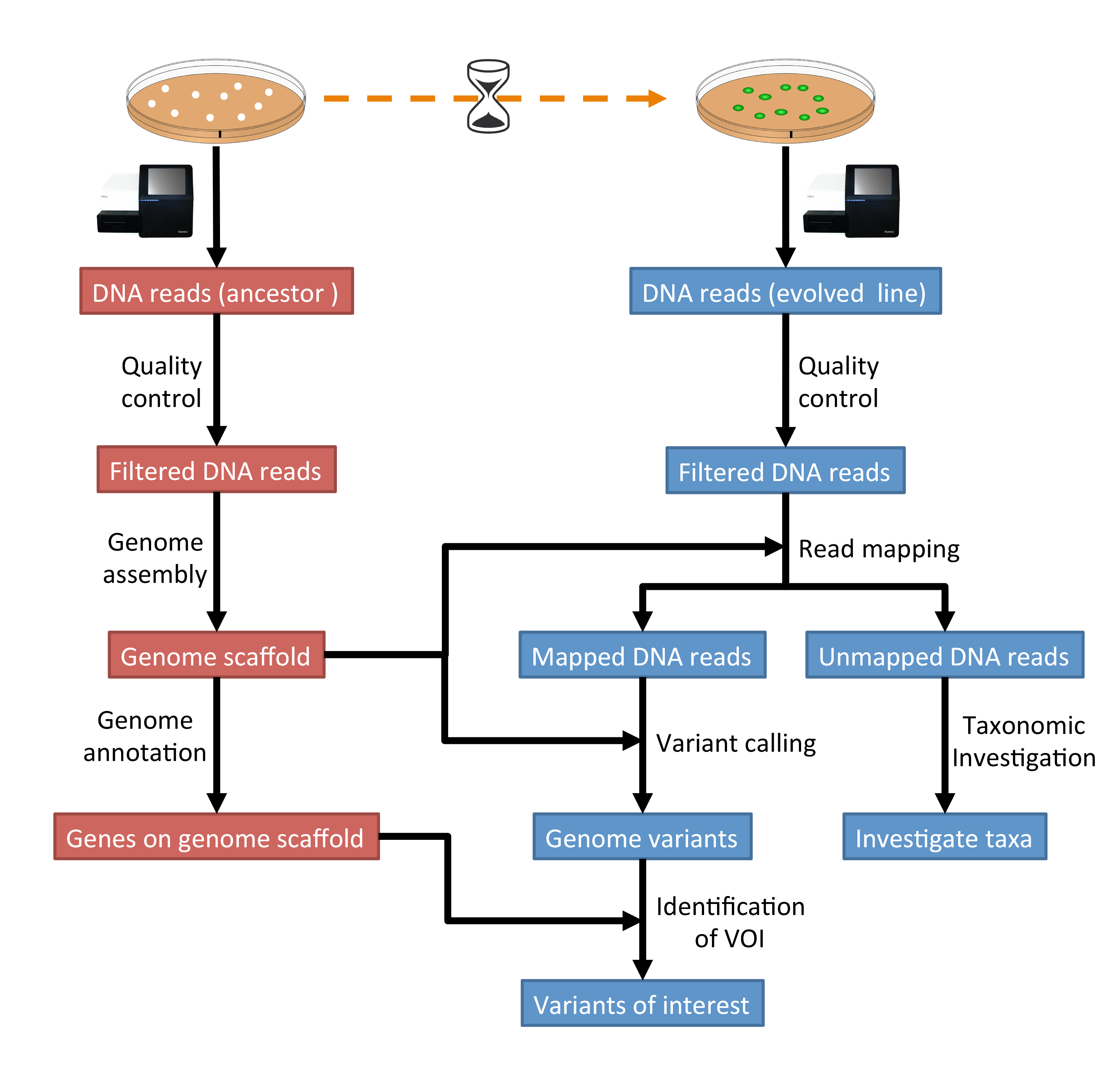
This is an introductory tutorial for learning computational genomics mostly on the Linux command-line. You will learn how to analyse next-generation sequencing (NGS) data. The data you will be using is real research data. The final aim is to identify genome variations in evolved lines of bacteria that can explain the observed biological phenotypes. Until 2020, Sebastian was teaching this material in the Massey University course Genome Science.
More information about other bioinformatics material and our past research can be found on the former webpages of the Schmeier Group (https://www.schmeierlab.com).
Note
A PDF-version of this tutorial can be downloaded here.